| World Journal of Oncology, ISSN 1920-4531 print, 1920-454X online, Open Access |
| Article copyright, the authors; Journal compilation copyright, World J Oncol and Elmer Press Inc |
| Journal website http://www.wjon.org |
Letter to the Editor
Volume 9, Number 2, April 2018, pages 66-67
Concomitant ATM Mutations Identified by Next Generation Sequencing in a Patient With New-Onset Acute Myeloid Leukemia Following Imatinib Treatment for Chronic Myeloid Leukemia
Matthew K. Steina, b, d, Melissa Crawleya, Eric Vickc, Mike G. Martina, b
aDepartment of Hematology/Oncology, West Cancer Center, Memphis, TN, USA
bDepartment of Internal Medicine, University of Tennessee Health Science Center, Memphis, TN, USA
cCollege of Medicine, University of Tennessee Health Science Center, Memphis, TN, USA
dCorresponding Author: Matthew K. Stein, Department of Internal Medicine, University of Tennessee Health Science Center, 956 Court Ave., Suite H314, Memphis, TN 38163, USA
Manuscript submitted March 11, 2018, accepted March 27, 2018
Short title: Dual ATM Mutations With New AML
doi: https://doi.org/10.14740/wjon1094w
| To the Editor | ▴Top |
ATM mutations have been described in breast, lung, hematologic, gastrointestinal and genitourinary malignancies [1, 2]. Pritchard et al report an 11.8% incidence of pathogenic germline mutations in DNA-repair genes among metastatic prostate cancer patients [2]. Of 20 genes surveyed, all associated with hereditary cancer predisposition syndromes, ATM variants comprised 13% (11/84). Further, a “second “hit” somatic aberration” was found in 36/61 (59%) with available tumor sequencing data.
We report a 70-year-old man diagnosed with new-onset acute myeloid leukemia (AML) following 14-year maintenance with imatinib for chronic-phase chronic myeloid leukemia (CML) with complete molecular remission and undetectable BCR/ABL transcripts. Cytogenetic analysis from initial bone marrow biopsy demonstrated karyotype 46,XY,del(7)(q22) with fluorescence in situ hybridization negative for BCR/ABL, t(8;21) and t(15;17). Next generation sequencing (NGS) (FoundationOne Heme panel, Foundation Medicine) described four pathogenic mutations: ATM (W579*), DNMT3A (R882H), IDH1 (R132C) and NPM1 (W288fs*10+). Variants of unknown significance (VUS) were found in six genes including ATM (G2863V), which in silico analysis utilizing PolyPhen-2 software (http://genetics.bwh.harvard.edu/pph2/) predicted as probably damaging (Table 1) [3]. Bone marrow biopsy following induction and salvage chemotherapy with cytarabine and idarubicin showed residual disease. During induction, gallbladder adenocarcinoma was diagnosed by biopsy. The patient subsequently elected for hospice care. During treatment, his family declined genetic counseling.
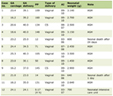 Click to view | Table 1. In Silico Analysis of VUS Identified by NGS of Bone-Marrow Cells in a CML Patient with BCR/ABL-negative AML |
One reality of NGS is the prospect of identifying pathogenic germline variants in cancer susceptibility genes [4]. Determining the significance of these mutations is difficult when NGS does not utilize matched-normal comparison DNA. Additionally, the clinical significance of VUSs identified by NGS remains elusive [5].
Following NGS, our CML patient was diagnosed with BCR/ABL-negative AML with concomitant single nucleotide variants in ATM: a pathogenic, non-sense mutation disrupting the protein kinase domain (W579*) and VUS point mutation (G2863V). Public databases including ClinVar and dbSNP supported the pathogenicity of W579*. A search for both variants on ExAc returned no results. Coupled with in silico analysis predicting G2863V as likely pathogenic, his AML harbored two presumed pathogenic mutations in ATM, although germline status was never established.
This case illustrates both the tremendous value and unique challenge of interpreting genomic information supplied by NGS. Although rare, BCR/ABL-negative AML has been previously reported in CML patients following imatinib therapy [6-8], with one CML cohort identifying 3/1,701 patients with development of AML or high-risk myelodysplastic syndrome (MDS) with eventual AML transformation [6]. In our patient, we postulate the ATM lesions as contributing to susceptibility to AML following imatinib treatment. Additionally, the identification of coexistent mutations in ATM underscores a need for access to genetic counseling and clinical germline testing. Utilized in genetic counseling, we also propose in silico analysis as a clinical tool for oncologists to better characterize VUS supplied by NGS.
Author Contributions
MM and MS designed the research. MS analyzed data. MS, MM, MC and EV wrote the paper.
Conflict of Interest
The authors have no conflict of interest to disclose.
| References | ▴Top |
- Cremona CA, Behrens A. ATM signalling and cancer. Oncogene. 2014;33(26):3351-3360.
doi pubmed - Pritchard CC, Mateo J, Walsh MF, De Sarkar N, Abida W, Beltran H, Garofalo A, et al. Inherited DNA-repair gene mutations in men with metastatic prostate cancer. N Engl J Med. 2016;375(5):443-453.
doi pubmed - Hahn AW, Giri S, Patel D, Sluder H, Vanderwalde A, Martin MG. Next-generation sequencing and in silico analysis facilitate prolonged response to pazopanib in a patient with metastatic urothelial carcinoma of the renal pelvis. J Natl Compr Canc Netw. 2015;13(10):1181-1185.
doi pubmed - Schrader KA, Cheng DT, Joseph V, Prasad M, Walsh M, Zehir A, Ni A, et al. Germline variants in targeted tumor sequencing using matched normal DNA. JAMA Oncol. 2016;2(1):104-111.
doi pubmed - Hahn AW, Martin MG. Precision medicine: lessons learned from the SHIVA trial. Lancet Oncol. 2015;16(16):e580-581.
doi - Kovitz C, Kantarjian H, Garcia-Manero G, Abruzzo LV, Cortes J. Myelodysplastic syndromes and acute leukemia developing after imatinib mesylate therapy for chronic myeloid leukemia. Blood. 2006;108(8):2811-2813.
doi pubmed - Krysiak K, Christopher MJ, Skidmore ZL, Demeter RT, Magrini V, Kunisaki J, O’Laughlin M, et al. A genomic analysis of Philadelphia chromosome-negative AML arising in patients with CML. Blood Cancer J. 2016;6:e413.
doi pubmed - Perel JM, McCarthy C, Walker O, Irving I, Williams B, Kennedy GA. Clinical significance of development of Philadelphia-chromosome negative clones in patients with chronic myeloid leukemia treated with imatinib mesylate. Haematologica. 2005;90(Suppl):ECR25.
pubmed
This article is distributed under the terms of the Creative Commons Attribution Non-Commercial 4.0 International License, which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
World Journal of Oncology is published by Elmer Press Inc.


